Our next guest post is by David Baltrus (@surt_lab) on his group’s preprint Extensive Phenotypic Changes Associated with Large-scale Horizontal Gene Transfer, posted on bioRviv here.
The function of modern pickup trucks is usually to haul heavy loads from point A to point B. However, the F-150 sitting in my driveway right now looks very different from its Model T ancestor from ~100 years ago. Over the years, as truck design has been modified and improved, all of the parts (brakes, air conditioning systems, doors, wheels, etc…) have been crafted to fit and work efficiently together. In process, each of the parts you see on a pickup truck today have been selectively co-evolving with all of the other design elements on the truck. The function of a house is to provide shelter.You can easily extend the the co-evolutionary metaphor from above to explain how different aspects of the house I live in relate to one another.
Some time ago, someone had the brilliant idea merge houses and pickup trucks into a camper. These hybrids between pickups and houses provide the functionality of being able to drive around, while also maintaining the ability to provide shelter. However, in the beginning, these hybrids likely didn’t accelerate as fast and consumed more energy and resources than unweighted pickups. They were likely a little taller than unweighted pickups, and as such might not be able to use certain bridges or tunnels. The brakes probably didn’t work as well. I can go on and on, but that would belabor the point I’m trying to make. In the beginning, if you just place two independently designed systems together Rube Goldberg style, the result will likely be functional but inefficient. Over the years, as engineers have worked to smoothly merge all of the systems of pickup and house together, campers have gotten much better at doing both jobs simultaneously.
I’ll freely admit that the above two paragraphs are a bit of a reach as a metaphor, but I think it provides a nice tangible context of what we are trying to measure within this preprint. Think of two independently evolving microbial lineages (lineage A = pickup, lineage B = house). Each lineage is really good at performing one function that the other can’t. For the sake of example, we’ll say that lineage A can perform phenotype A (catabolism of naphthalene) while lineage B can perform phenotype B (resistance to antibiotic). Horizontal gene transfer (HGT), the movement of genes and pathways between lineages without reproduction, is a major evolutionary force within microbes and has pretty much affected evolutionary dynamics of all life at one time or another. Through HGT, the lineage A is able to acquire the pathways encoding phenotype B, and in doing so is now able to invade a previously unoccupied niche. Presumably moth balls drenched in antibiotic is the main food source available in this niche. Microbial evolution at it’s finest. Although we don’t directly explore these phenotypes within our system, they are both influenced by HGT (napthalene and antibiotic resistance).
However, the genome of lineage A has not previously co-evolved with the pathway encoding phenotype B. Immediately at the moment of transfer, it’s very likely that the pathway encoding phenotype B will not work efficiently within the new genomic context of lineage A. Just like inefficiencies of design with the first hybrid camper, there will be costs and side effects as genetic and regulatory networks are merged.Lots of papers on this including here, here, here, and here, as well as an overview I wrote for TREE this year. For instance, the production of additional proteins requires extra resources in terms of nucleotides and amino acids. The pathway for phenotype B may not be regulated properly, and proteins from pathway B may fold incorrectly leading to cytotoxic effects. There are a lot of potential inefficiencies that we as researchers can think of immediately after merging a new pathways into an independent genome through HGT. These costs and side effects have been widely demonstrated throughout systems and transfer events. If you’ve ever had to deal with cloning a “toxic” protein into E. coli, you’ve dealt with costs of HGT. Sure you might be able to get the construct in, but the cells die if you express too much protein. In industry these costs are known as metabolic burden because they lower the total yield of the product of interest. I’m certainly not the first to point out what happens when you glue together systems that have been independently evolving. However, much of the previous research into costs of HGT deals with measuring single phenotypes (like growth rate) to quantify costs. However, to bring back the camper metaphor, this is like only measuring only how acceleration time is affected by gluing a house to a pickup. Even though there are many other aspects (miles per gallon, height, which roads you can take) that could affected, we don’t really have an appreciation or baseline for the variety of ways that HGT can screw up cell systems. Many of these previous experiments have also only dealt with relatively small genomic fragments (<40kb) moving around (extending the metaphor further, this is like setting up a tent in the back of a pickup). We really don’t generally know much about how HGT of larger fragments affects cells and if everything learned previously scales up with size of transfer.
Although I’ve been interested in HGT as an evolutionary process since grad school, my postdoc work was focused more on genomics and the evolution of virulence. However, as a postdoc I discovered that one strain of Pseudomonas syringae had acquired a 1Mb megaplasmid compared to a hundred or so other strains we sequenced. As one of those experiments you never think will work, but which are worth a try, I placed an antibiotic marker onto this megaplasmid, mixed strains together, and waited for the magic to happen. Turns out, strictly through comparative genomics and some hunches, that we’ve found the largest self-transmissible plasmid known to date within bacteria. Since Pseudomonas genomes are typically 4-6Mb, adding an extra Mb through HGT means that cells can increase their genome size by ~20% overnight and add 700 or so novel ORFs in the process. In other words, a nice bookend to studies of costs of HGT using relatively small fragments.
In a second paper currently unpublished, we have shown that transfer of this megaplasmid to other P. syringae or P. stutzeri strains entails about a 20% fitness cost as measured by competitive assays. In this preprint we further explore what other kinds of costs and secondary effects occur with acquisition of the megaplasmid by P. stutzeri. For instance, we’ve been able to show that acquisition of this megaplasmid sensitizes strains to certain classes of antibiotics like naldixic acid and ciprofloxin as well as DNA intercalating dyes like 7-hydroxycoumarin (Fig. 2). These results pretty much came from fishing…we send paired strains off to Biolog (either containing or lacking the megaplasmid) hoping that we might discover that the megaplasmid imparted some beneficial effect or new niche. Instead, we found hypersensitivity to antibiotics…and made delicious lemonade out of lemons.

Fig. 2 HGT of a megaplasmid lowers resistance to nalidixic acid. Megaplasmid containing strain in grey, wild type strain in black
The next result we found was that megaplasmid acquisition sensitizes strains to heat (Fig. 3). Again, this was the produce of experimental stumbling. Although I was able to conjugate the megaplasmid into P. stutzeri, my technician was not. We compared notes and found that I was plating post conjugation cultures at 27 degrees while Kevin was plating at 37 degrees. Turns out that acquisition of the megaplasmid makes 37 degrees a lethal temperature for P. stutzeri.

Fig. 3 HGT of a megaplasmid sensitizes P. stutzeri to high (ish) temperatures. Serial dilutions were plated out at 27 degrees and 37 degrees. White circle represents where bacteria plated out but didn’t grow.
The last couple of results (increases in motility, decreases in biofilm formation, sensitivity to other bacterial toxins) were really the result of random observations we had and followed up on or shots in the dark. For motility, we just decided to test whether motility was altered because it was fairly straightforward. For the biofilm result, we noticed that older cultures of bacteria spooled around a pipette tip which we used to inoculate the cultures but that megaplasmid containing strains didn’t have this behavior. The bacteriocin result was discovered by my grad student Brian Smith as he was trying to mate the megaplasmid into Pseudomonas aeruginosa (and was failing). Although all of these results have been witnessed independently in other bacteria in other contexts, demonstrating that one HGT event can change all of these phenotypes is pretty unprecedented as far as I can tell. I’m exciting to figure out what the basis of these changes are and if they’re independent. Spoiler, currently thinking it’s due to interference with some stress responses and gene regulation.
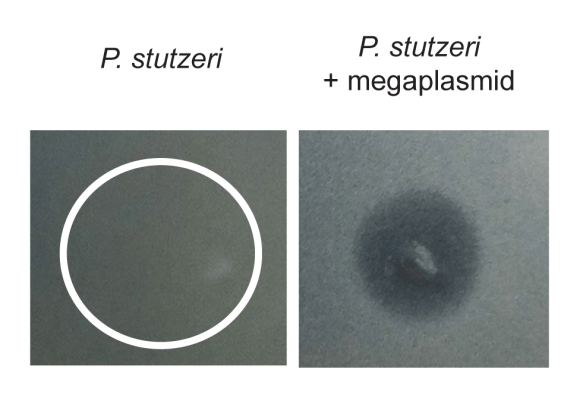
Fig. 4 HGT of a megaplasmid sensitizes P. stutzeri to killing by P. aeruginosa supernatant. On left is wild type P. stutzeri on right is megaplasmid containing strain spotted with supernatant. Lack of bacterial growth is only seen on right only where supernatant spotted (darker part).
So there you have it. Placed within the camper metaphor that I started out with, we’ve discovered that placing a house into a pickup bed not only changes acceleration but has a bunch of other previously unrecognized changes. The engine now requires a higher octane of gas to work, the engine overheats much more easily, etc…Given my evolutionary background, I’m very interested in figuring out how acquisition of the megaplasmid changes future evolutionary dynamics within these bacterial populations. Can my little camper still travel on all the same roads as the pickup (does acquisition of the megaplasmid change evolvability or evolutionary trajectory)? How does amelioration of these fitness costs change systems level relationships between bacterial phenotypes? Hopefully I’ll have another preprint up in the next year addressing some of these questions!


For your possible interest, I have a webpage with >800 references about HGT at panspermia.org/virus.htm. For the references click “What’sNEW”