The following post is by Joe Pickrell [@joe_pickrell] on his preprint Joint analysis of functional genomic data and genome-wide association studies of 18 human traits, available on bioRxiv here.
Until recently, the field of human genetics struggled to identify genetic variants that influence complex traits and diseases like height or diabetes. With the arrival of genome-wide association studies (GWAS), studies now regularly identify tens to hundreds of genomic regions that contain such variants. The question going forward is clear: how do these variants influence traits?
One way to answer this question involves annotating variants according to their potential functions–does a given variant change the sequence of a protein? Or does it disrupt the splicing of a gene? Or does it fall in a regulatory region in an important cell type? Many groups (like those that are part of the ENCODE project) are generating hundreds of datasets that are potentially informative about these types of questions. But which of these hundreds of datasets are relevant when studying a given trait?
In this preprint, I develop a statistical method (an empirical Bayes hierarchical model) that takes summary statistics from a genome-wide association study of a given trait and identifies the types of genomic annotation that are relevant for the trait; software implementing this method is available here. I then applied this method to a set of 18 traits and 450 genomic annotations. Feedback on the method itself is of course welcome, but I’d also like to highlight what I think are the most interesting biological results:
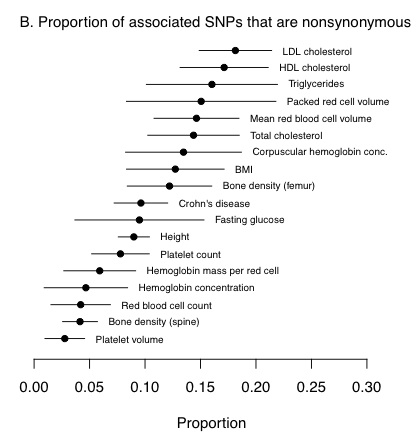
- The relative importance of protein-coding versus regulatory variants varies across traits. The fraction of GWAS hits driven by changes in protein sequence depends on the trait, ranging from a low of around 2% up to around 20% (see above).
- Repressed chromatin is depleted for loci that influence traits. I was surprised to find that the most informative type of information for interpreting a GWAS is often repressed chromatin, which is depleted for loci influencing traits. This type of chromatin covers up to 70% of the genome.
- Cell type-specific DNase-I hypersensitive sites are enriched for loci that influence traits. A “hypothesis-free” scan across all regulatory regions in many tissues can identify unexpected connections between traits and tissues. For example, loci that influence bone density are enriched in gene regulatory regions in muscle tissue, and loci that influence Crohn’s disease are enriched in regulatory regions in fibroblasts.
- Incorporating functional information into a GWAS increases power to detect loci. Finally, re-weighting a GWAS using this method increases the number of loci identified in each GWAS by around 5%; many of the loci identified with this method have been replicated in larger studies.
I’m hopeful that this method (and others like it) will be useful in making the transition from identifying statistical associations in a GWAS to understanding the underlying biology; comments and criticisms are welcome.

Pingback: Most viewed on Haldane’s Sieve: December 2013 | Haldane's Sieve
Might be fun to also consider doing the proportion of variance explained attributable to different functional categories. Or perhaps that is too much of a headache to work out due to difficulties of not really knowing which SNP is true association (nor its effect size).
Can you do that from summary statisitcs? I think you may need to get to the genotype / phenotype data to calculate the variance explained properly.
Pingback: Author post: VSEAMS: A pipeline for variant set enrichment analysis using summary GWAS data identifies IKZF3, BATF and ESRRA as key transcription factors in type 1 diabetes | Haldane's Sieve